
Research Summary
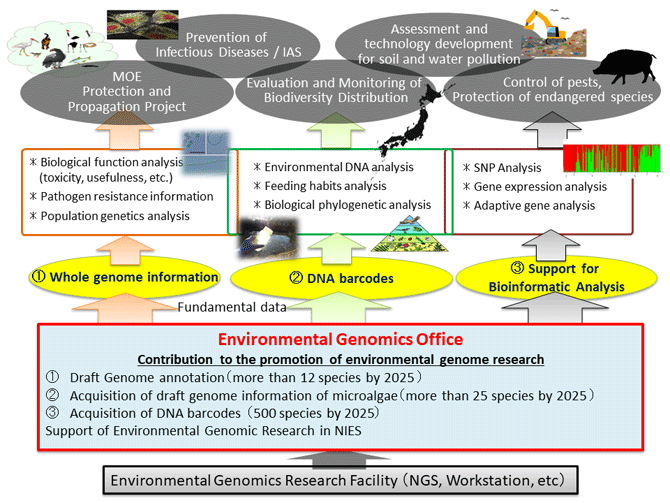
In biodiversity research using genetic information, the quality and quantity of fundamental genetic information have a significant impact on the resolution and reliability of the analysis results. In this project, we will mainly undertake (1) acquisition of whole genome information of wildlife, and (2) collection of nucleotide sequence information of DNA regions suitable for species discrimination of organisms (DNA barcodes). This genetic information provides useful data that can be used immediately in studies of wildlife conservation and propagation, extermination, and pest distribution. It will also contribute to the accumulation of reference data that will accelerate biodiversity assessment techniques using DNA obtained from environmental samples such as soil and water (environmental DNA), thereby improving the accuracy of biodiversity surveys in environmental impact assessments and monitoring. In addition, we will assist in research planning, experiments, and analysis techniques for environmental DNA analysis and the acquisition of genome-wide intra- and interspecies variation information.
Research Objectives
We will provide genetic information that forms the basis for the fifth five-year plan of NIES and related research in Japan and overseas, and contribute to the promotion of various advanced analyses.
Acquisition and release of draft genome information and DNA barcodes
For species subject to Wild Bird and Animal Protection and Propagation Projects, we will provide whole genome information with annotation information indicating the location and function of genes as basic data for improving the efficiency of protection and propagation projects and for analysis to determine the causes of population decline. For microalgae such as blue-green algae, we will continuously acquire draft genome information to understand the genetic diversity in Japan. This is expected to provide distribution information with academic novelty and functional information, such as toxicity and usefulness. To improve the accuracy of biodiversity surveys using environmental DNA, we will collect and enrich DNA barcode information of organisms living in Japan.
Support of genome-related research in NIES
We assist in genome-related research within the institute by obtaining information on genome-wide mutations such as single nucleotide polymorphisms, supporting environmental DNA analysis, and contributing to the analysis of wildlife populations using information on their genetic structure and distribution.
Research Staff
NAKAJIMA Nobuyoshi, KONDO Natsuko, ANDO Haruko、KATAYAMA Masafumi, ISHII Yumiko
Contact address
Environmental Genomics Office NAKAJIMA Nobuyoshi
TEL:029-850-2490
Mail:naka-320 (Please attach "@ nies.go.jp" to the end)
Related link
Genome Database
Environmental Specimen Time Capsule Building

